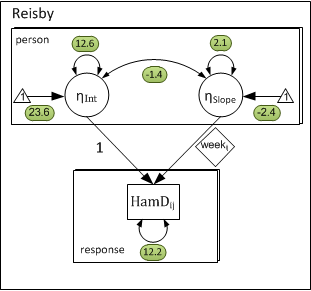
The model equations and matrices in the present example are identical to the previous. However, there are two key features that make the present model distinct: (1) level-1 observations represent reaction times nested within individuals and (2) the level-1 predictor measures time since the study began (measurement occasion), leading to a latent growth curve model (LGC). We have chosen to structure the data in a manner consistent with the mixed-effects modeling approach to LGC analysis, which also allows us to draw an explicit parallel between the present model and the more general random-slopes model (example 2) in which the level-1 predictor represents another attribute of the level-1 unit. The data for the present example were drawn from the Reisby et al. (1977) example described in Hedeker’s (2004) introduction to growth modeling chapter. The outcome is ratings on the Hamilton depression rating scale (Hamilton, 1960). Depressions scores were taken over a period of weeks. A baseline was taken (week 0). Ratings were then taken after a week of the subjects consuming a placebo (week 1) and the following four weeks (week 1-5) participant took a depression drug.
Table of Contents
Model Equations
Level-1
\[ HamD_{ij}= 1_{ij} \times \eta_{Intj} + week_{ij} \times \eta_{Slopej} + e_{ij} \]
Level-2
\[ \eta_{Intj} = \gamma_{00} + u_{0j} \] \[ \eta_{Slopej} = \gamma_{10} + u_{1j} \]
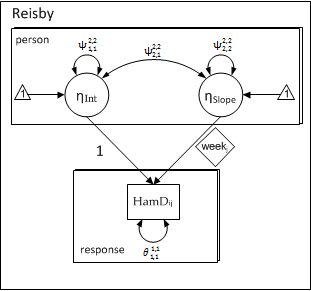
As in the previous model, the coefficients are latent variables and the predictors (\( 1_{ij} \) & \( week_{ij} \) are fixed. The following path-diagram has a one-to-one correspondence to the first and second level equations for a random-slopes model.
The above model is identical to the random-slopes model presented earlier:
\[ y_{ij}^1 = 1_{ij}^{1,2} \times \eta_{1j}^2 + week_{ij}^{1,2} \times \eta_{2j}^2 + e_{ij}^1, \]
\[ \eta_{1}^2 = \alpha_{1}^2 + \zeta_{1j}^2 \] \[ \eta_{2}^2 = \alpha_{2}^2 + \zeta_{2j}^2 \]
Path diagram
xxM Model Matrices
Level-1: Within matrices
Residual Covariance Matrix
As before, we have a single dependent variable and hence a single parameter level-1 residual variance \( (\theta_{1,1}) \) at level-1. Hence, the residual covariance or theta matrix is a (1×1) matrix:
\Theta^{1,1}=
\begin{bmatrix}
\theta^{1,1}_{1,1} \\
\end{bmatrix}
\]
Level-2: Within Matrices
At level-2, we have two latent variables: intercept and slope. Hence, we have two latent means and a covariance matrix.
Latent Means
The latent variable mean matrix is a (2×1) matrix:
\[
\alpha^2=
\begin{bmatrix}
\alpha^2_1 \\
\alpha_2^2
\end{bmatrix}
\]
\( (\alpha_1^2) \) is the mean of the intercept and \( (\alpha_2^2) \) is the mean of the slope parameter or the average effect of \( week_{ij} \) on \( HamD_{ij} \), a measure of depression.
Latent Factor Covariance Matrix
The latent covariance matrix is a (2×2) matrix with two variances and single covariance:
\[
\Psi^{2,2}=
\begin{bmatrix}
\psi^{2,2}_{1,1} \\
\psi^{2,2}_{2,1} & \psi^{2,2}_{2,2}
\end{bmatrix}
\]
\( \psi_{1,1}^{2,2} \) is the variance of the intercept factor representing variability in the intercept of \( y_{ij} \) across persons and \( \psi_{2,2}^{2,2} \) is the variance of the slope parameter representing between-persons variability in the effect of \( week_{ij} \) on \( HamD_{ij} \). Finally, \( \psi_{2,1}^{2,2} \) is the covariance between the intercept and slope factors.
Across level matrices: Person to Response
As described above, we need to capture the effect of level-2 intercept and slope factors on the level-1 dependent variable using a factor-loading matrix with fixed parameters.
Factor-loading matrix
The factor-loading matrix \( (\Lambda^{1,2}) \) has a single row and two columns (1×2):
\[
\Lambda^{1,2}=
\begin{bmatrix}
1.0 & week_{ij}
\end{bmatrix}
\]
The first column is fixed to 1.0, whereas the second column is fixed to person-specific value of \( week_{ij} \). Collectively, the within and across-level matrices described here specify all of the parameters necessary to model individual differences in over-time trajectories of reaction times across persons. Next, we provide SAS, MPlus, and xxM code for fitting LGCs.
Code Listing
xxM
As in the previous example, there is a single level-1 dependent variable and a single level-1 independent variable. The number of matrices is the same as before, and their dimensions are also identical. :
1. We do not wish to estimate any factor-loadings. Factor-loadings are fixed to 1.0 and \( week_{ij} \). Hence, both elements of pattern matrix are zero:\[ \Lambda^{pat}= \begin{bmatrix} 0.0 & 0.0 \end{bmatrix} \]
- We want to fix the first-factor loading to 1.0 (intercept). We use the value matrix to provide the fixed-value of 1.0 for the first factor-loading. The second factor-loading does not have a single fixed value for every observation. Instead each observation \( (i) \) would have its own value for that factor-loading \( (x_{ij}) \). Clearly, the value matrix cannot be used for providing individual specific fixed values. Hence, the second element in the value matrix is left as 0.0. xxM ignores it internally.\[ \Lambda^{val}= \begin{bmatrix} 1.0 & 0.0 \end{bmatrix} \]
- The job of fixing the factor-loading is left to the label matrix. A label matrix is used to assign labels to each parameter within the matrix. Label matrices can be used to impose equality constraints across matrices. Any two parameters with the same label are constrained to be equal. Label matrices are also used for specifying that a specific parameter is to be fixed to data-values. In this case, the first-label is irrelevant as that parameter has already been fixed to 1.0. We use a descriptive label \( lambda_{11} \) as the first label (something such as Justin would have worked as well). The second factor-loading is the one we are interested in. We want to fix the second factor-loading to the observation specific values of the predictor \( (week_{ij}) \). This is accomplished by using a two-part label: levelName.predictorName. In this case, the predictor is a response level variable. Hence, the first part of the label is response. The second part is the actual predictor name, in this case week. \[ \Lambda^{label}= \begin{bmatrix} l_{1,1} & week_{ij} \end{bmatrix} \]
lambda_label <- matrix(c("lambda_11", "response.week"), 2, 1)
The complete listing of xxM code for the latent growth curve (long version) example follows:
Load xxM and data
Construct R-matrices
For each parameter matrix, construct three related matrices:
- pattern matrix: A matrix indicating free or fixed parameters.
- value matrix: with start or fixed values for corresponding parameters.
- label matrix: with user friendly label for each parameter. label matrix is optional.
# Within-level matrices Response: level-1 matrices Theta : observed residual
# covariance
th1_pat <- matrix(1, 1, 1)
th1_val <- matrix(20, 1, 1)
# Subject: level-2 matrices Psi : Laent growth factor covariance
ps2_pat <- matrix(1, 2, 2)
ps2_val <- diag(0.1, 2)
# Psi :: Laent growth factor means
al2_pat <- matrix(1, 2, 1)
al2_val <- matrix(c(25, -1), 2, 1)
# Across-level matrices Subject to respons (level-2 to level-1) factor
# loading matrix
ly12_pat <- matrix(0, 1, 2)
ly12_val <- matrix(c(1, 0), 1, 2)
ly12_label <- matrix(c("one", "response.week"), 1, 2)
Construct main model object
xxmModel() is used to declare level names. The function returns a model object that is passed as a parameter to subsequent stattements.
lgc <- xxmModel(c("response", "subject"))
## A new model was created.
Add submodels to the model objects
For each declared level xxmSubmodel() is invoked to add corresponding submodel to the model object. The function adds three pieces of information:
1. parents declares a list of parents of the current level.
2. variables declares names of observed dependent (ys), observed independent (xs) and latent variables (etas) for the level.
3. data R data object for the current level.
lgc <- xxmSubmodel(model = lgc, level = "response", parents = "subject", ys = "depression",
xs = "week", etas = , data = response)
## Submodel for level `response` was created.
lgc <- xxmSubmodel(model = lgc, level = "subject", parents = , ys = , xs = ,
etas = c("intercept", "week_slope"), data = subject)
## Submodel for level `subject` was created.
Add Within-level parameter matrices for each submodel
For each declared level xxmWithinMatrix() is used to add within-level parameter matrices. For each parameter matrix, the function adds the three matrices constructed earlier:
- pattern
- value
- label (optional)
lgc <- xxmWithinMatrix(lgc, "response", "theta", th1_pat, th1_val)
##
## 'theta' matrix does not exist and will be added.
## Added `theta` matrix.
lgc <- xxmWithinMatrix(lgc, "subject", "psi", ps2_pat, ps2_val)
##
## 'psi' matrix does not exist and will be added.
## Added `psi` matrix.
lgc <- xxmWithinMatrix(lgc, "subject", "alpha", al2_pat, al2_val)
##
## 'alpha' matrix does not exist and will be added.
## Added `alpha` matrix.
Add Across-level parameter matrices to the model
Pairs of levels that share parent-child relationship have regression relationships. xxmBetweenMatrix() is used to add corresponding parameter matrices connecting the two levels.
- Level with the independent variable is the parent level.
- Level with the dependent variable is the child level.
For each parameter matrix, the function adds the three matrices constructed earlier:
- pattern
- value
- label (optional)
lgc <- xxmBetweenMatrix(lgc, "subject", "response", "lambda", ly12_pat, ly12_val,
ly12_label)
##
## 'lambda' matrix does not exist and will be added.
## Added `lambda` matrix.
Estimate model parameters
Estimation process is initiated by xxmRun(). If all goes well, a quick printed summary of results is produced.
lgc <- xxmRun(lgc)
## ------------------------------------------------------------------------------
## Estimating model parameters
## ------------------------------------------------------------------------------
## 2663.4456544893
## 2619.5947785561
## 2580.1443385522
## 2546.2979080330
## 2517.4873994193
## 2493.5468495741
## 2476.1883025592
## 2465.3974417533
## 2462.9357470790
## 2460.5754518934
## 2457.0606447791
## 2424.5113353373
## 2370.6011886285
## 2348.5078782042
## 2336.9817970649
## 2316.6849583854
## 2311.5507070718
## 2310.2782595431
## 2260.9062913709
## 2249.6717647161
## 2249.4574873888
## 2248.5781901713
## 2246.9823608581
## 2245.0177518635
## 2243.6138389961
## 2240.6089389811
## 2239.2298214422
## 2238.0561513054
## 2234.4571065836
## 2230.5787416242
## 2227.9193579953
## 2223.5371518942
## 2220.2383182681
## 2219.5309099732
## 2219.1104671881
## 2219.0531076585
## 2219.0396765010
## 2219.0376169346
## 2219.0375139502
## 2219.0375117402
## 2219.0375116516
## Model converged normally
## nParms: 6
## ------------------------------------------------------------------------------
## *
## 1: response_theta_1_1 :: 12.217 [ 0.000, 0.000]
##
## 2: subject_psi_1_1 :: 12.629 [ 0.000, 0.000]
##
## 3: subject_psi_1_2 :: -1.421 [ 0.000, 0.000]
##
## 4: subject_psi_2_2 :: 2.079 [ 0.000, 0.000]
##
## 5: subject_alpha_1_1 :: 23.577 [ 0.000, 0.000]
##
## 6: subject_alpha_2_1 :: -2.377 [ 0.000, 0.000]
##
## ------------------------------------------------------------------------------
Estimate profile-likelihood confidence intervals
Once parameters are estimated, confidence inetrvals are estimated by invoking xxmCI() . Depending on the the number of observations and the complexity of the dependence structure xxmCI() may take very long. xxMCI() displays a table of parameter estimates and CIS.
View results
A summary of results may be retrived as an R list by a call to xxmSummary()
xxmSummary(lgc)
## $fit
## $fit$deviance
## [1] 2219
##
## $fit$nParameters
## [1] 6
##
## $fit$nObservations
## [1] 375
##
## $fit$aic
## [1] 2231
##
## $fit$bic
## [1] 2255
##
##
## $estimates
## child parent to from label estimate
## 1 response response depression depression response_theta_1_1 12.217
## 4 subject subject intercept intercept subject_psi_1_1 12.629
## 5 subject subject intercept week_slope subject_psi_1_2 -1.421
## 6 subject subject week_slope week_slope subject_psi_2_2 2.079
## 7 subject subject intercept One subject_alpha_1_1 23.577
## 8 subject subject week_slope One subject_alpha_2_1 -2.377
## low high
## 1 10.263 14.7007
## 4 6.954 21.3955
## 5 -3.919 0.3632
## 6 1.252 3.3661
## 7 22.492 24.6627
## 8 -2.792 -1.9619
Free model object
xxM model object may hog a large amount of RAM outside of R’s memory. This memory will automatically be released, when R’s workspace is cleared by a call to rm(list=ls()) or at the end of the R session. Alternatively, xxmFree() may be called to release memory.
lgc <- xxmFree(lgc)
rm(list = ls())
Proc Mixed
Proc Mixed data = riesby covtest;
CLASS subject;
MODEL HamD = week/s;
RANDOM Intercept week/subject = subject G type = UN;
RUN;
SAS code for a random-slopes model uses a CLASS statement to identify the level-2 units, in this case person. The MODEL statement estimates the fixed-effects \( (\alpha) \). The RANDOM statement specifies that the level-1 intercepts and the effect of level-1 predictor \( week_{ij} \) is allowed to vary across “subject” (i.e., persons). The covariance among the random-effects (G) is freely estimated (specified by “type = UN”). The G matrix corresponds to the xxM \( \psi \) matrix. Finally, like all regression models, Proc Mixed estimates the residual variance of the level-1 dependent variable \( (\theta_{1,1}) \) by default. The important thing to note is that there is one-to-one correspondence between the parameters estimated in Proc Mixed and SEM.
For the current dataset, the parameter estimates are: